Day 229: Coronavirus modeling – Part 2
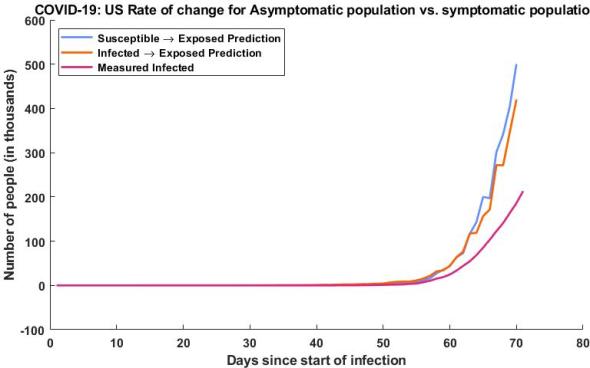
Here we have my estimated exposed population vs the measured infected population, the exposed population is calculated two ways, the first using our susceptible population dynamics (blue) and the second is using our measured infected rate (orange). This is compared to our measured infected since we would expect to see a much higher exposed (asymptomatic) population than infected (symptomatic) population, which we do.
I’ve made a lot of progress! Unfortunately, my model needs some more work and the code is pretty messy right now. For now, I’ll share some of my outputs and discuss what I need to do to finish my assignment. It turns out I have a few extra days to finish the work, I thought it was due Monday, instead it is due Wednesday. The slides and write up are going to take the longest so I’m still crunched for time even though I am mostly done. Let’s go over it.
Modeling is basically a lot of assumptions. We use past values to predict future values. Thankfully there are a lot of people who came before me that did the hard work, they tracked pandemics and created the formula we still use to this day. There are things you can do differently to solve the set of equations that make up your model. There are also ways to add things, remove things, etc. For example, my flu pandemic model was a “very simple” method for predicting the number of people who would test positive for the flu based on previous values.
Notice I said test positive, this is because we don’t know how many people out there had the flu that didn’t get tested, but we can use the previous values of people tested to predict the future values. In my case, my model was a stochastic model based on several simplifying assumptions that would’ve been publishable about 10 years ago. In other words, it had already been done, I was just sort of exploring someone else’s work. It was a good experience, but this time I wanted to step up my game so to speak.
Enter the SEIR model. As I mentioned yesterday, this model has 4 states Susceptible, Exposed, Infected, and Recovered. We estimate the rates that people move from one state using a set of ordinary differential equations (ODE). The set of equations are old, (think early 1900’s) so it’s not like I’m doing anything groundbreaking using them here. In fact, we use this type of model all the time.
There are two ways to use it, there is the deterministic way (with no randomness), this simplifies the model and makes the ODE easier to work with. By that I mean solve for the variables that make up ODE. So there are a handful of variables depending on the formulation and you can add more depending on how fancy you want to be with your prediction. The catch is, the simpler your model the more assumptions you need, but the more complex your model the more variables you have to solve for and that often requires several simplifying assumptions.
The second way to use this is in a stochastic model, this model adds noise you your measurement. There are advantages and disadvantages to using this model type as well. The biggest issue is the ODE are a pain to solve for, so you need to use techniques that to anyone who doesn’t use them on a regular basis look super convoluted. Math is hard… However, the good news is that for large N (population size) the models converge! So for large populations (like say the size of the US for example) we can use the deterministic model.
I did not.
I am using a simplified stochastic model, I’m doing this because I want my model to be applicable for both large populations and small populations. If I’m going to put the work in, I want it to be as useful as possible damn it. I’m also doing some semi-novel things with how I make my predictions. I’ll break it down in my video that I need to make for class, so I’ll share that here. In the meantime, I need to finish updating my model and make some predictions for a week or two ahead then I need to do my write up and make my slides.
Yesterday I used the infected measurement to predict the death rate. We saw that it worked out pretty well (even though I only predicted a day ahead, which as I mentioned isn’t too hard to do). This means my variables are in a good place to approximate the data. However, we don’t need to go in just one direction, we can go backwards too. I can use the death rate to determine the infection rate. I’ve also calculated the population for the exposed case as well, so we can predict infected two different ways and compare them to see how the model is working.
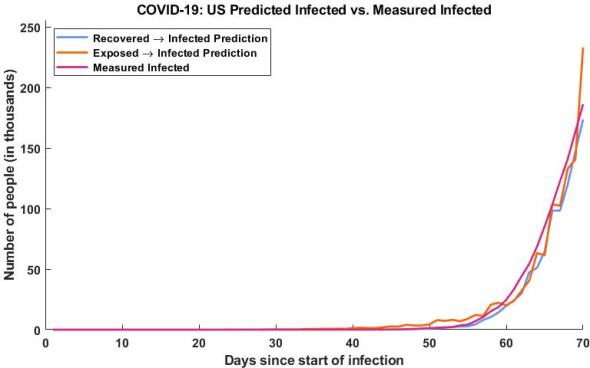
Determining the infection rate using the recovered population estimate (blue), the infection rate using our exposed population estimate (orange), and the actual measured infection rate (purple) This is not a prediction based on a measured value, it is a prediction based on a different prediction this is much different than trying to predict our infected rate based on our measured infected rate.
As you can see, the model is off to a good start, something of interest to note, the exposed and recovered populations are the estimates I created using this model, so we are back calculating our infected rate. This means two things, the first is that we are no longer making a prediction based on collected data, this is once removed (a prediction from another prediction). The second is that we can very accurately recover our infected state from our other estimated states. While this is useful to validate our model, it isn’t particularly useful for say a hospital to determine the needed equipment for the following week. It does however mean that I am close to finishing my model and with a few changes I can make predictions based on previous values.
With that, it’s back to work I go.



But enough about us, what about you?