‘Virtual virus’ unfolds the flu on a CPU
The flu virus can be pretty nasty — it’s quick to evolve — which means yearly flu shots are needed and then it’s only a guess to which strain will be the most prevalent. Well new research aims to change all that, by combining experimental data from X-ray crystallography, NMR spectroscopy, cryoelectron microscopy and lipidomics (the study of cellular lipid networks), researchers have built a complete model of the outer envelope of an influenza A virion for the first time. So would that make it a computer virus, virus?
The approach, known as a coarse-grained molecular dynamics simulation, has allowed them to generate trajectories at different temperatures and lipid compositions – revealing various characteristics about the membrane components that may help scientists better understand how the virus survives in the wild or find new ways to combat it.
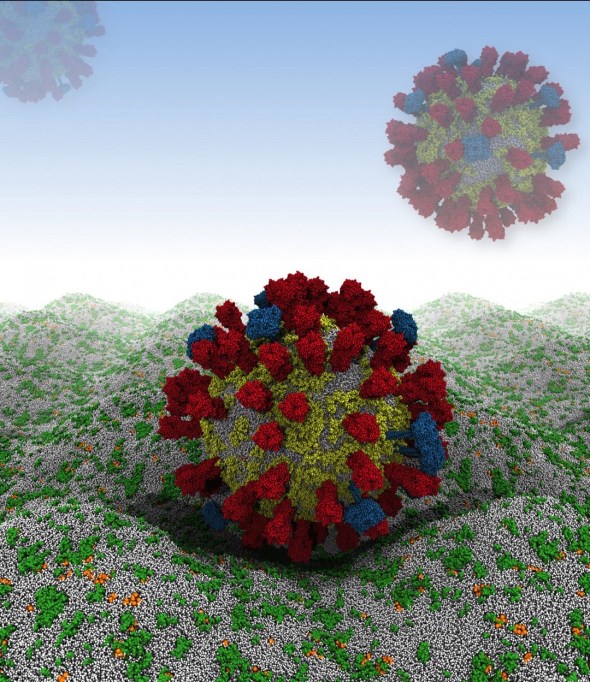
Their computer simulation begins by rendering the virus as a relatively large, 73-nanometer ball of loosely packed lipids. This ball then relaxes down into a smaller, 59-nanometer virion within 300 nanoseconds – an imperceptible amount of time on the macroscopic level, but roughly 1/15th of the simulation’s total run time. The viral spike proteins are then embedded into the lipid envelope individually, before adding solvent to the system.
“In the current publication, this is just a single virion in a water droplet — but what could really get interesting is if we start putting in other molecules of interest, such as therapeutic agents,” said Tyler Reddy, a postdoctoral fellow at the University of Oxford.
From the simulation, the team has found that the viral spike proteins protruding from the virion’s membrane spread out, rather than aggregating close together. This is key to the strength of the interactions between influenza A virions and host cells, which are determined by the number of spike proteins that can engage with receptors.
“If the separation of the spike proteins is compatible with the ‘arms’ of Y-shaped, bivalent antibodies, this information might be exploited in therapeutic design, so that two antigens may be bound simultaneously for enhanced association,” Reddy said.
The work builds on that of Daniel Parton, at the time, Reddy was working with molecular dynamics simulations for a membrane protein system relevant to achondroplasia, the most common cause of human dwarfism. He was impressed by Parton’s plan to construct and simulate the lipid envelope of the entire flu virion, and later decided to take aim at it himself.
The researchers also observed that the Forssman glycolipid had a role in preventing protein aggregation and slowing down protein diffusion. To which the team indicated it would be important to include glycolipids in future virion simulations given their influence on the biophysical properties observed. The extended sugar head groups of glycolipids may also mask antibody accessibility of the M2 proton channels in the flu envelope — the target of commonly prescribed anti-influenza drugs based on adamantane derivatives.
Understanding the membrane envelope’s structural dynamics also provides insight into the wide-ranging survival times of the virion in different environments, such as fresh-water rivers. Previous studies have indicated that the presence of influenza A in rivers has allowed waterfowl to be simultaneously exposed to source flu strains and residual anti-viral compounds excreted by local human populations, potentially giving rise to drug-resistant influenza strains. The simulation currently monitors the virion’s stability on the micro-second scale, and it will be a challenge to assess stability over much longer time scales.
“We are a long way from being able to perform molecular dynamics simulations that span the year time scale,” Reddy said.
“Nonetheless, we now have a platform for looking at influenza A virion behavior in silico, and perhaps certain compounds or solutions could be used to accelerate destabilization on an observable time scale,” Reddy said.
“We’re making the coordinate data freely available in the hopes that other groups have interesting ideas for use with this model as well, and so that they can criticize and help improve the model.”
The work is a huge leap towards finding a way to stop the virus, or at the very least find new ways to combat it. It’s exciting to think how fast the work has progressed and how much faster it will progress as computers become faster and technologies to explore the unseen become better.
Sources:
Reddy, T., Shorthouse, D., Parton, D., Jefferys, E., Fowler, P., Chavent, M., Baaden, M., & Sansom, M. (2015). Nothing to Sneeze at: A Full-Scale Computational Model of the Human Influenza Virion Biophysical Journal, 108 (2) DOI: 10.1016/j.bpj.2014.11.195



But enough about us, what about you?